Dictionary Update
After creating a Dictionary you can update the terms and synonyms from several different databases (using their flat-files) or the internal Biowarehouse database. This operation allows the user to add contents to a dictionary, which can come from several sources.
Loading a Dictionary to the clipboard:
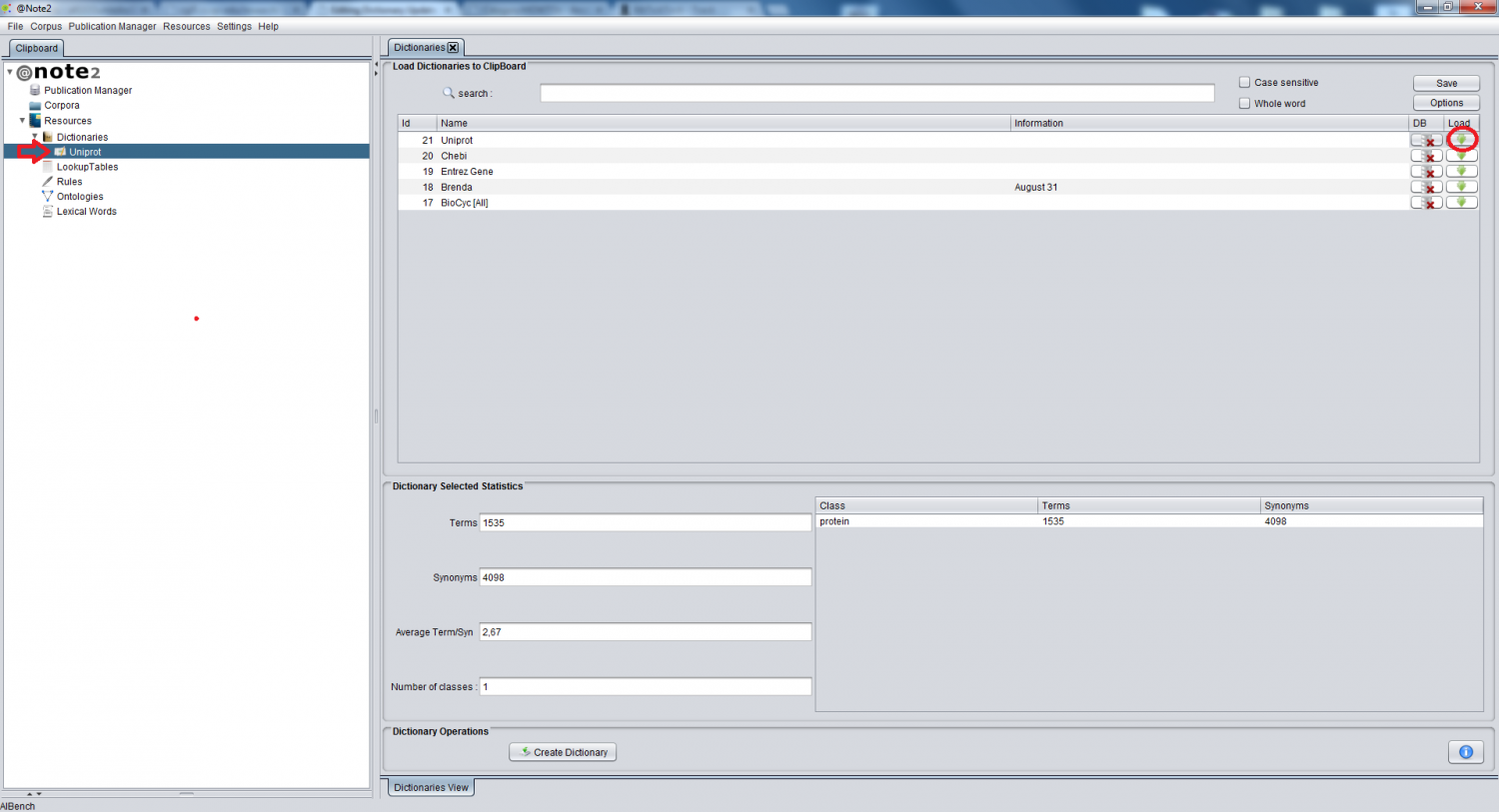
To have access to dictionary management operations, you have to firstly load the dictionary datatype to the clipboard. To do this, the Dictionaries datatype on the clipboard needs to be selected and a view is launched in the right hand side panel containing the list of available dictionaries. To load a Dictionary to the clipboard, select the appropriate row on the table and click in the load icon (rightmost button, highlighted by a red circle). The dictionary is now in the clipboard (highlighted by a red arrow) being possible to perform operations over it.
Updating Dictionary
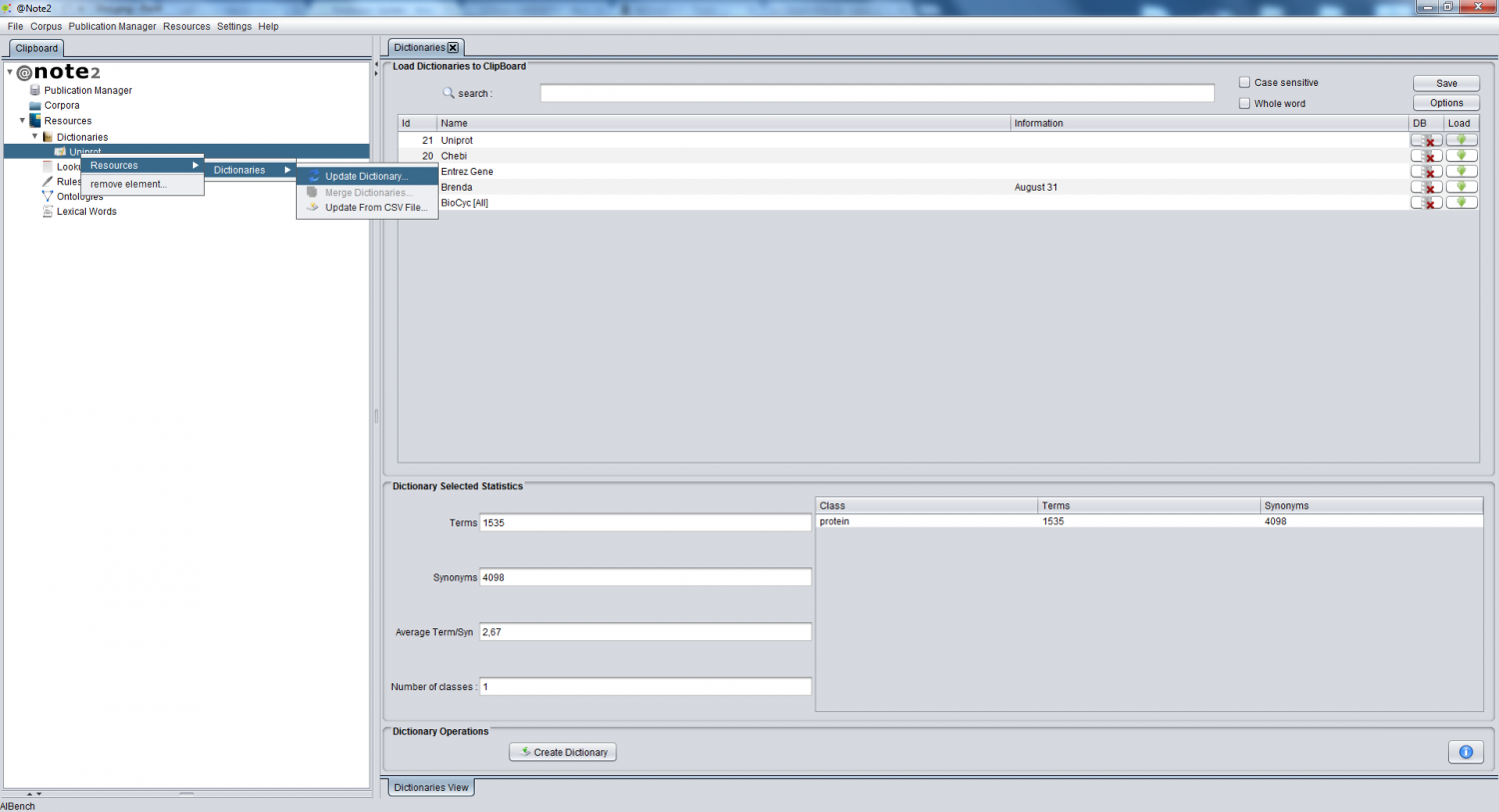
To update the dictionary contents, right click over the desired dictionary (highlighted by a red arrow in the image above), selecting the option Dictionary Update.
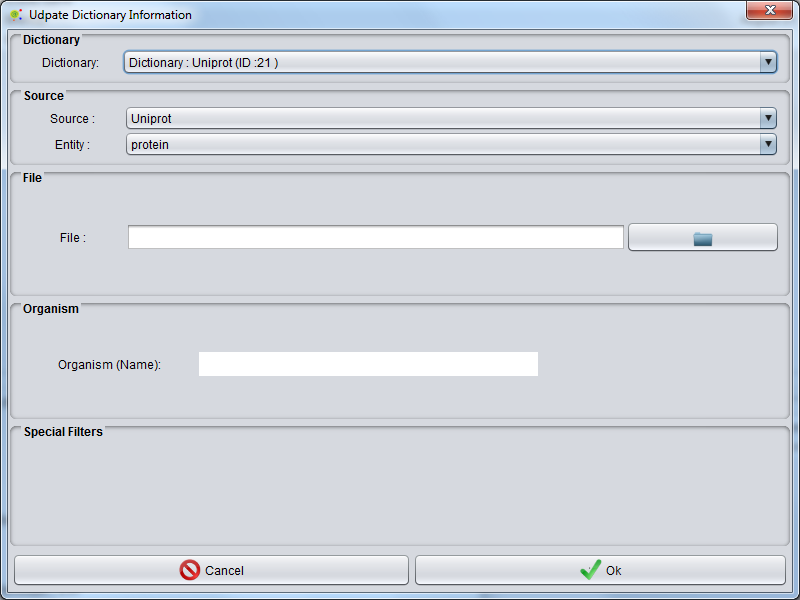
A GUI is launched where you can select the source of the dictionary, the entity class for the terms to be uploaded and choose the file to upload.
Some sources allow selecting the organism.
Flat Files Supported:
Currently, @Note supports the following sources:
- BioCyc flatfiles (http://biocyc.org/download.shtml) ;
- ChEBI names.tsv flatfiles (ftp://ftp.ebi.ac.uk/pub/databases/chebi/Flat_file_tab_delimited/names.tsv.gz);
- NCBI Taxonomy names.dmp flatfiles (ftp://ftp.ncbi.nih.gov/pub/taxonomy/taxdmp.zip);
- UniProtKB/Swiss-Prot flatfiles (ftp://ftp.uniprot.org/pub/databases/uniprot/current_release/knowledgebase/complete/uniprot_sprot.dat.gz);
- NCBI Gene flatfiles (ftp://ftp.ncbi.nlm.nih.gov/gene/DATA/GENE_INFO).
- Brenda old flat files (2012)
- Kegg old flat files (2012)
- Drugbank drugbank vocabulary tsv file (https://www.drugbank.ca/releases/latest#open-data).
After this operation, you can view a resources update report