Difference between revisions of "Dictionary Update"
From Anote2Wiki
Anote2Wiki (talk | contribs) |
Anote2Wiki (talk | contribs) |
||
| Line 9: | Line 9: | ||
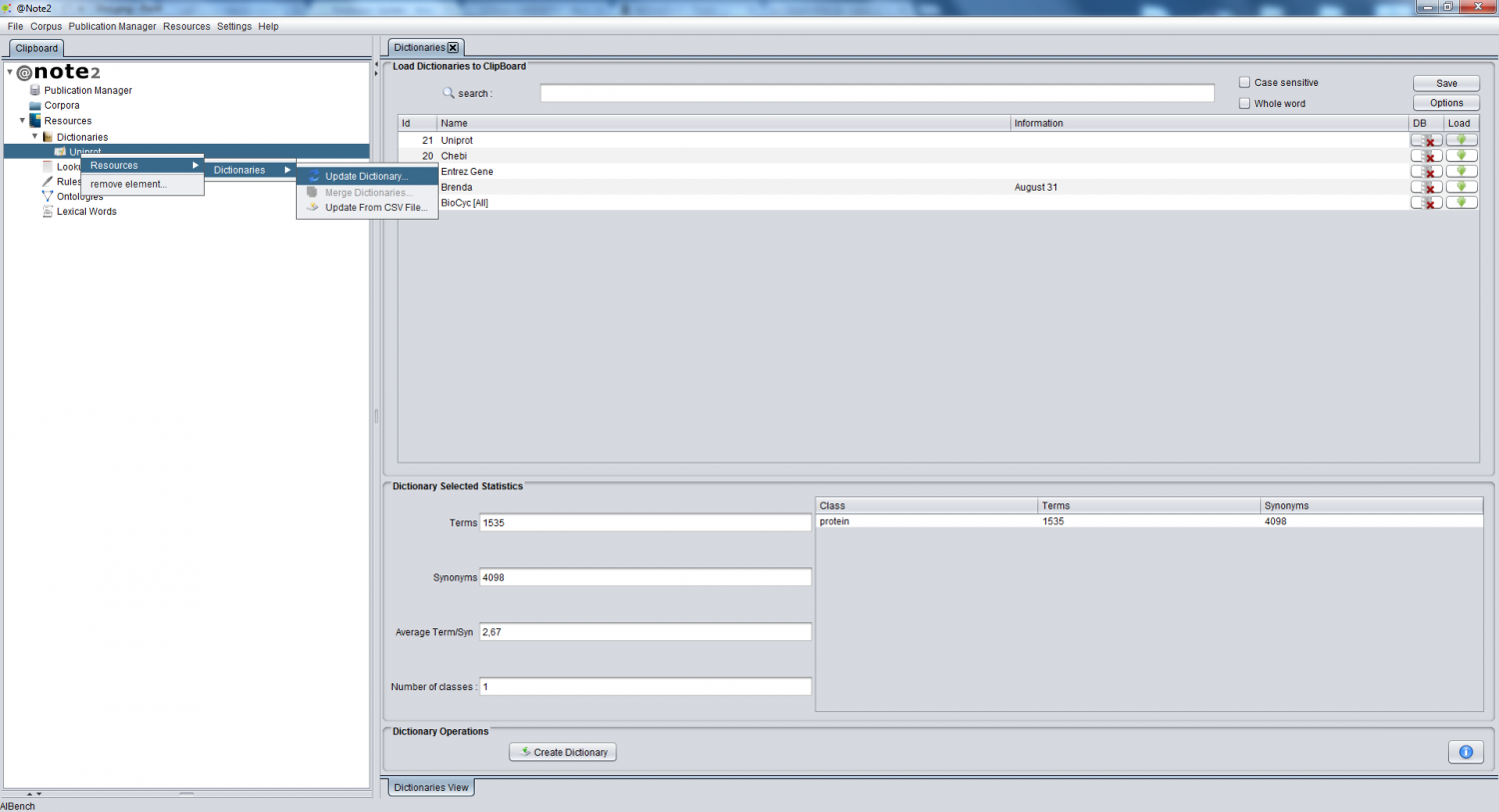
For have access to dictionary operations, first you have to load dictionary data-type to clipboard by selected dictionary bottom ( red circle ). Now in clipboard are available dictionary data-type (red arrow). | For have access to dictionary operations, first you have to load dictionary data-type to clipboard by selected dictionary bottom ( red circle ). Now in clipboard are available dictionary data-type (red arrow). | ||
| − | [[File:Dictionary_Update.png|center]] | + | [[File:Dictionary_Update.png|1500px|center]] |
== Update Dictionary: == | == Update Dictionary: == | ||
| Line 15: | Line 15: | ||
For update dictionary content press left mouse bottom on Dictionary data-type (image up). | For update dictionary content press left mouse bottom on Dictionary data-type (image up). | ||
| − | [[File:Dictionary_Update_GUI.png|center]] | + | [[File:Dictionary_Update_GUI.png|800px|center]] |
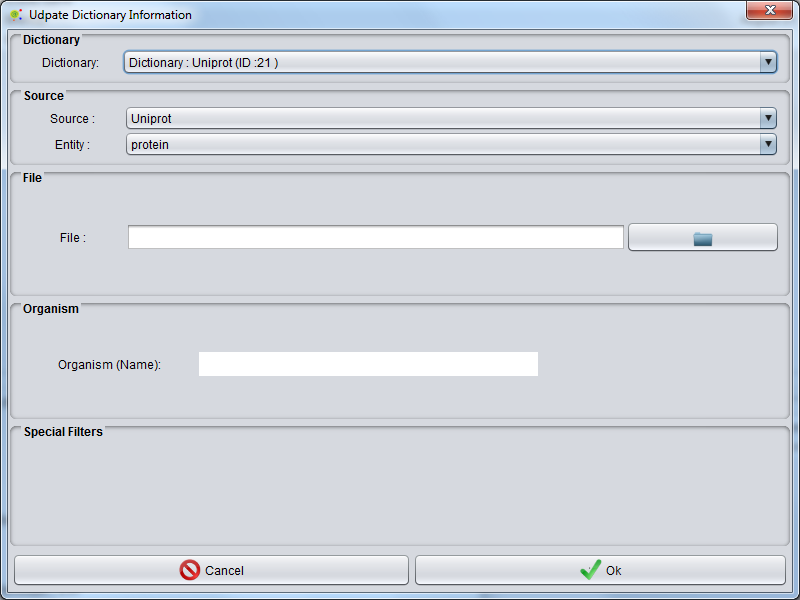
After appears Dictionary Update GUI and can select: | After appears Dictionary Update GUI and can select: | ||
Revision as of 09:25, 19 June 2012
After Create Dictionary you can update dictionary terms for some database flat-files or internal Biowarehouse database. This operation allows the user to add contents to a dictionary, which can come from several sources.
Load Dictionary to Clipboard:
For have access to dictionary operations, first you have to load dictionary data-type to clipboard by selected dictionary bottom ( red circle ). Now in clipboard are available dictionary data-type (red arrow).
Update Dictionary:
For update dictionary content press left mouse bottom on Dictionary data-type (image up).
After appears Dictionary Update GUI and can select:
- Database Flat Files type (see Flat Files Supported ).
- Entity Selected
- Flat File (Select files to access data)
- Organism - Some Flat File options than could select organism for entity selection
Flat Files Supported:
Currently, the system supports the following sources:
- BioWarehouse integrated databases (http://biowarehouse.ai.sri.com/);
- BioCyc flatfile(http://biocyc.org/download.shtml) ;
- ChEBI flatfiles (http://www.ebi.ac.uk/chebi/downloadsForward.do);
- NCBI Taxnonomy flatfiles (ftp://ftp.ncbi.nih.gov/pub/taxonomy/);
- UniProtKB/Swiss-Prot flatfiles (http://www.uniprot.org/downloads);
- MGI Entrez Genes flatfiles (ftp://ftp.informatics.jax.org/pub/reports/index.html).