Difference between revisions of "NER Model Creation"
Anote2Wiki (talk | contribs) (→Select Option) |
Anote2Wiki (talk | contribs) |
||
| (4 intermediate revisions by the same user not shown) | |||
| Line 12: | Line 12: | ||
== New Configuration or Load Configuration== | == New Configuration or Load Configuration== | ||
| − | A wizard will be presented to configure the model creation. The first step allows to select two options: Create New BioTML Model Configuration or Load BioTML Model Configuration. To start a new model | + | |
| + | A wizard will be presented to configure the model creation process. The first step allows to select two options: Create New BioTML Model Configuration or Load BioTML Model Configuration. | ||
| + | To start a new model configuration, select '''Create New BioTML Model Configuration''' and press the '''Next''' button.<br /> | ||
| Line 19: | Line 21: | ||
== Select NLP Tokenizer System == | == Select NLP Tokenizer System == | ||
| − | A GUI is presented to select the possible NLP systems that are integrated in the BioTML framework. Those systems are used to | + | |
| − | The possible NLP systems to be chosen are the ClearNLP, | + | A GUI is presented to select the possible NLP systems that are integrated in the BioTML framework. Those systems are used to perform the tokenization of all documents to create a data matrix for machine learning algorithms.<br /> |
| + | The possible NLP systems to be chosen are the ClearNLP, Stanford Core NLP and OpenNLP. Each system contains a description that is presented on this GUI. | ||
| Line 26: | Line 29: | ||
| − | == BioTML | + | == BioTML Feature Selection == |
| − | The features | + | |
| − | + | The selection of the features used by the machine learning algorithm as its training data matrix is done using the GUI below. | |
| + | Depending on the number and type of features selected, the produced model can fit the data with variable degree of accuracy. | ||
| + | The selection of the features can have a great impact in the predictive capability of the model, influencing precision, recall and accuracy of the model during the NER annotation.<br /><br /> | ||
| + | Warning: The number of features and some feature types can significantly increase the memory and CPU usage ! | ||
| Line 35: | Line 41: | ||
== BioTML Model Algorithm Configuration == | == BioTML Model Algorithm Configuration == | ||
| − | The machine learning algorithm is selected in this GUI. Advanced settings | + | |
| − | For further information | + | The machine learning algorithm to be used is selected in this GUI. |
| + | Currently available options are Conditional Random Fields (CRFs) implemented by the Mallet software and Support Vector Machines (SVMs) as implemented by LIBSVM software. | ||
| + | Advanced settings can appear depending on the selected algorithm type. <br /><br /> | ||
| + | For further information about those advanced configurations please visit the URLs: [http://mallet.cs.umass.edu/api/cc/mallet/fst/CRF.html CRF Informations] or [https://github.com/cjlin1/libsvm/blob/master/README SVM informations] | ||
| Line 42: | Line 51: | ||
| − | == | + | == Select NER Classes for ModelCreation == |
| + | |||
| + | |||
After the algorithm selection, a GUI is presented to select the possible NER classes that will be used to train the model. | After the algorithm selection, a GUI is presented to select the possible NER classes that will be used to train the model. | ||
| Line 49: | Line 60: | ||
| − | == | + | == Save Model File == |
| − | All configurations for NER Model creation are defined. In this | + | |
| + | All configurations for NER Model creation are now defined. In this GUI, you can define a folder or Zip file that will store the NER Model. | ||
Latest revision as of 14:06, 4 August 2015
Contents
Select Option
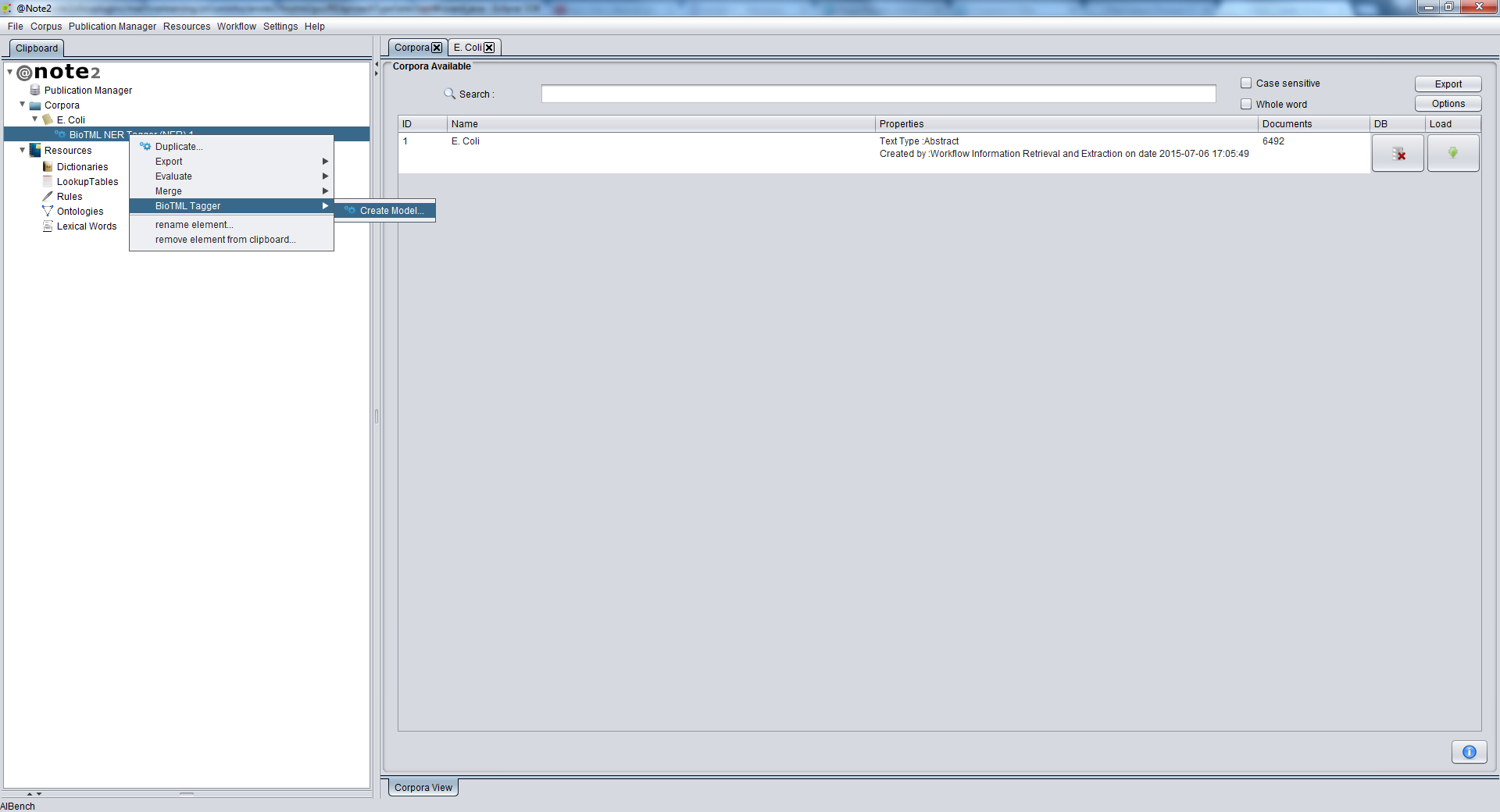
To create an NER Machine Learning Model using BioTML, the first step is to select the corpus with the NER annotations.
To do this, start by loading the desired NER Process to the Clipboard.
Selecting the NER Process, you should right click it and choose NER Process -> BioTML Tagger -> Create Model
New Configuration or Load Configuration
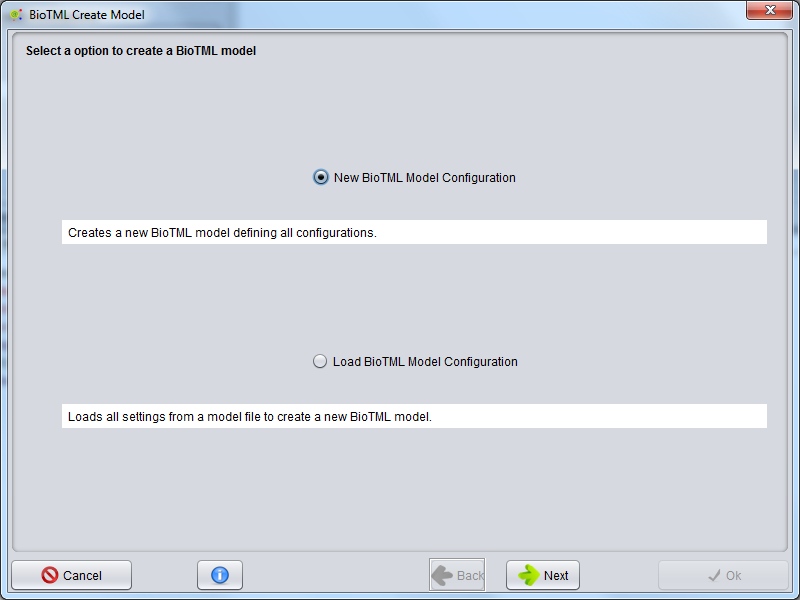
A wizard will be presented to configure the model creation process. The first step allows to select two options: Create New BioTML Model Configuration or Load BioTML Model Configuration.
To start a new model configuration, select Create New BioTML Model Configuration and press the Next button.
Select NLP Tokenizer System
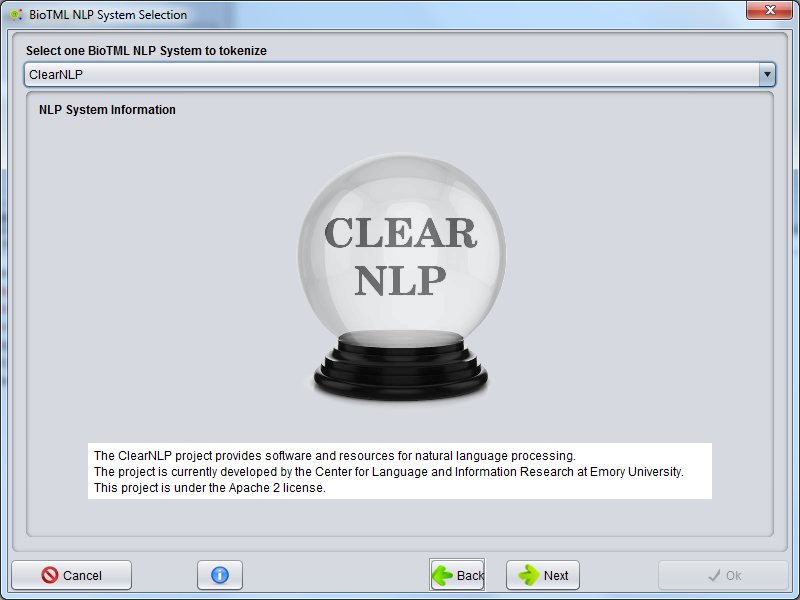
A GUI is presented to select the possible NLP systems that are integrated in the BioTML framework. Those systems are used to perform the tokenization of all documents to create a data matrix for machine learning algorithms.
The possible NLP systems to be chosen are the ClearNLP, Stanford Core NLP and OpenNLP. Each system contains a description that is presented on this GUI.
BioTML Feature Selection
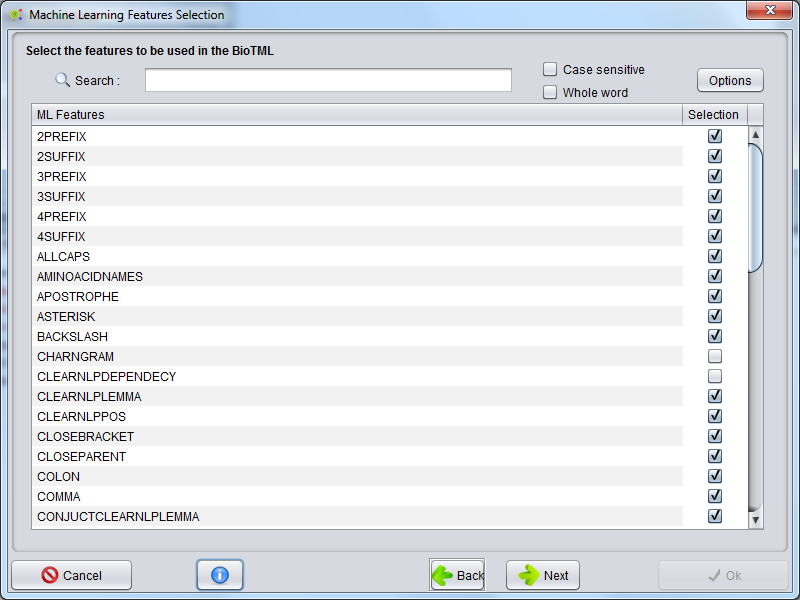
The selection of the features used by the machine learning algorithm as its training data matrix is done using the GUI below.
Depending on the number and type of features selected, the produced model can fit the data with variable degree of accuracy.
The selection of the features can have a great impact in the predictive capability of the model, influencing precision, recall and accuracy of the model during the NER annotation.
Warning: The number of features and some feature types can significantly increase the memory and CPU usage !
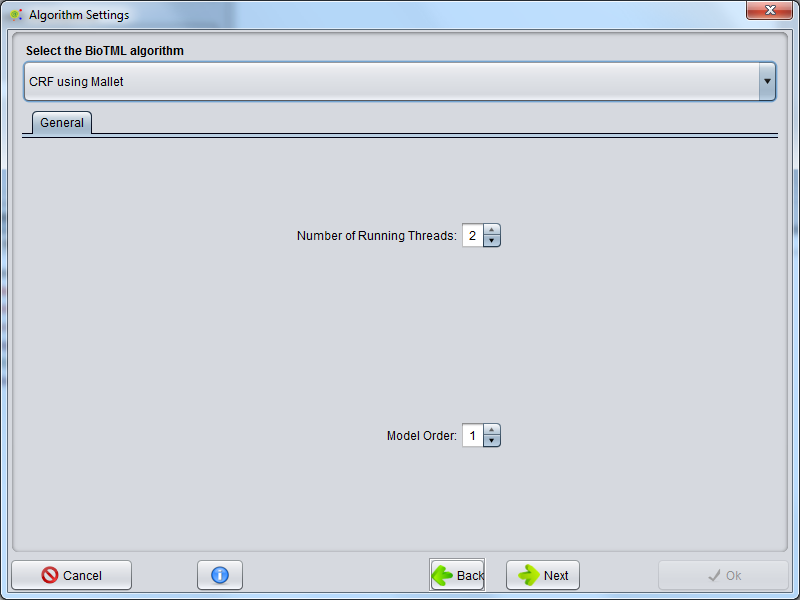
BioTML Model Algorithm Configuration
The machine learning algorithm to be used is selected in this GUI.
Currently available options are Conditional Random Fields (CRFs) implemented by the Mallet software and Support Vector Machines (SVMs) as implemented by LIBSVM software.
Advanced settings can appear depending on the selected algorithm type.
For further information about those advanced configurations please visit the URLs: CRF Informations or SVM informations
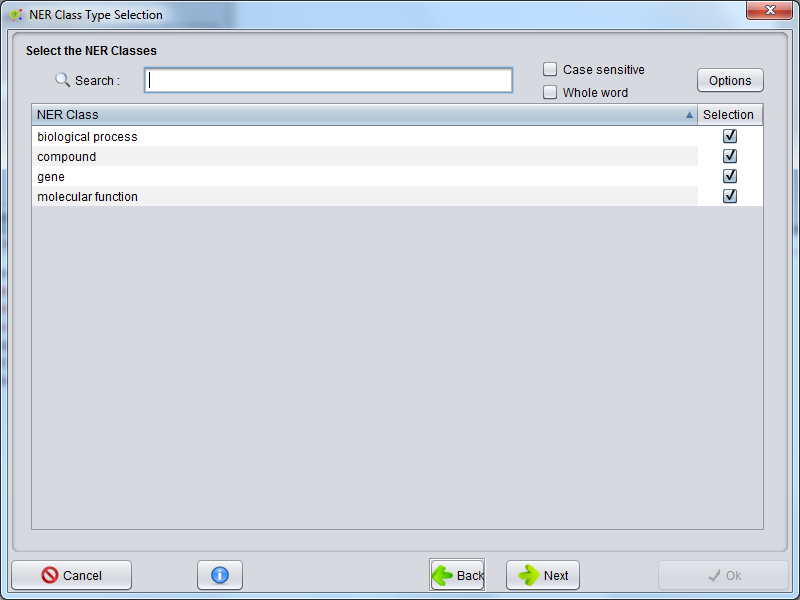
Select NER Classes for ModelCreation
After the algorithm selection, a GUI is presented to select the possible NER classes that will be used to train the model.
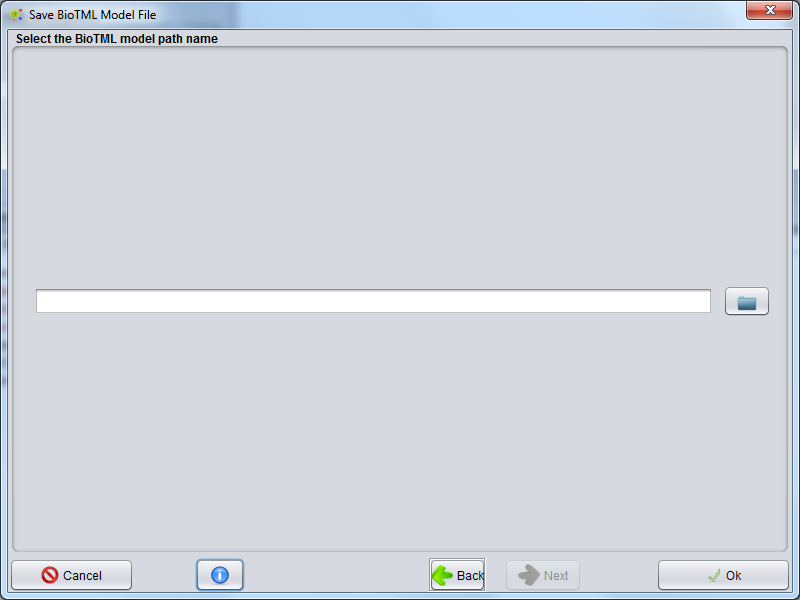
Save Model File
All configurations for NER Model creation are now defined. In this GUI, you can define a folder or Zip file that will store the NER Model.
Result
The NER model creation operation will start and a working window is shown, indicating the execution of the operation. The NER model creation operation will take a few minutes or hours, depending on corpus size and model configurations. When the process ends, a new Zip file containing the NER model will be added to the defined directory in the last GUI.