Corpus Create Annotation Schema By BioTML Tagger
Contents
Select Option
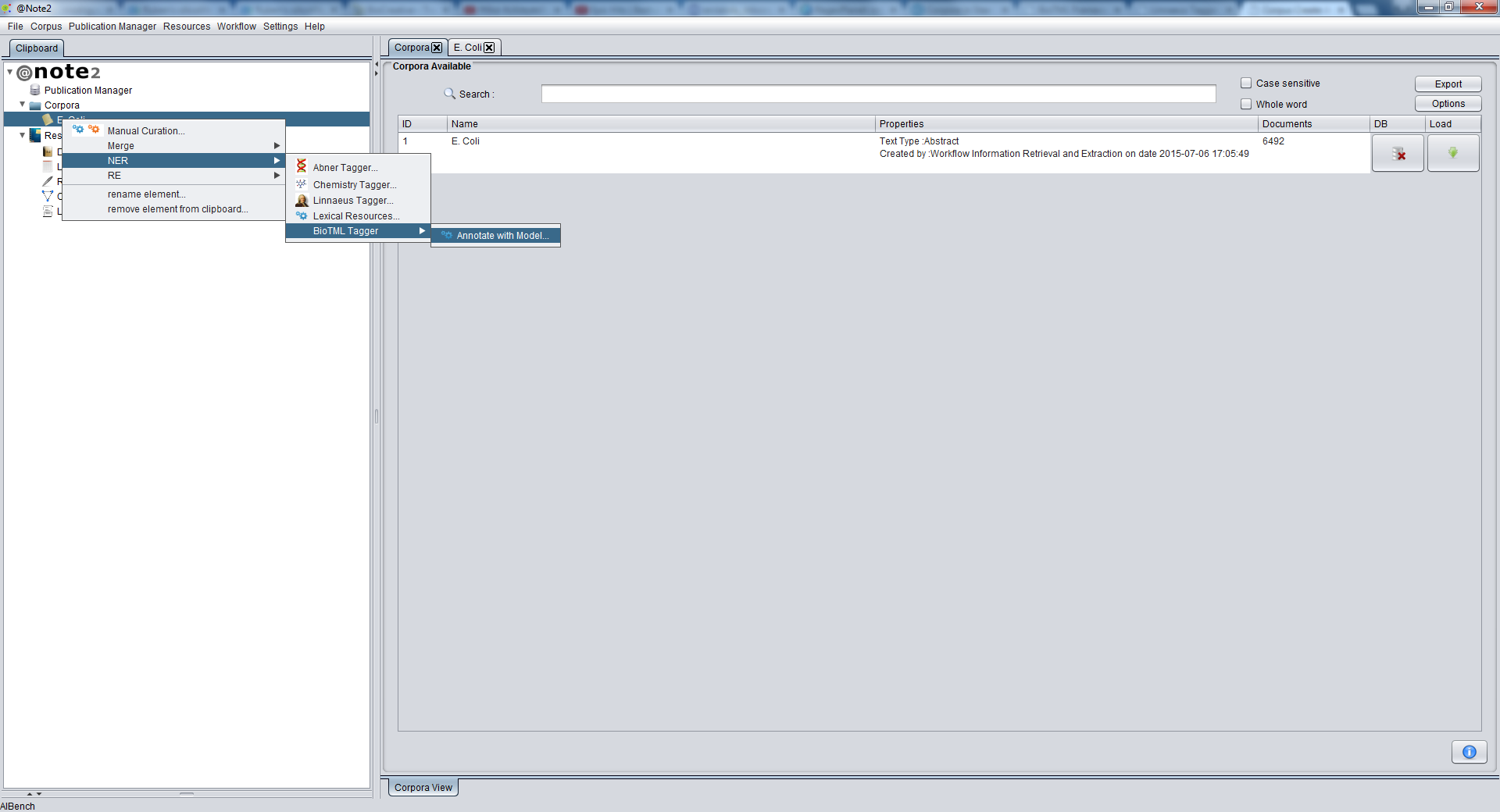
To perform a new NER (Entity Recognition) based in BioTML Tagger, start by loading a Corpus to the Clipboard.
Selecting the Corpus, you should right click over it and choose Corpus -> NER -> BioTML Tagger -> Annotate with Model
Load_BioTML_Model_File_For_Annotation
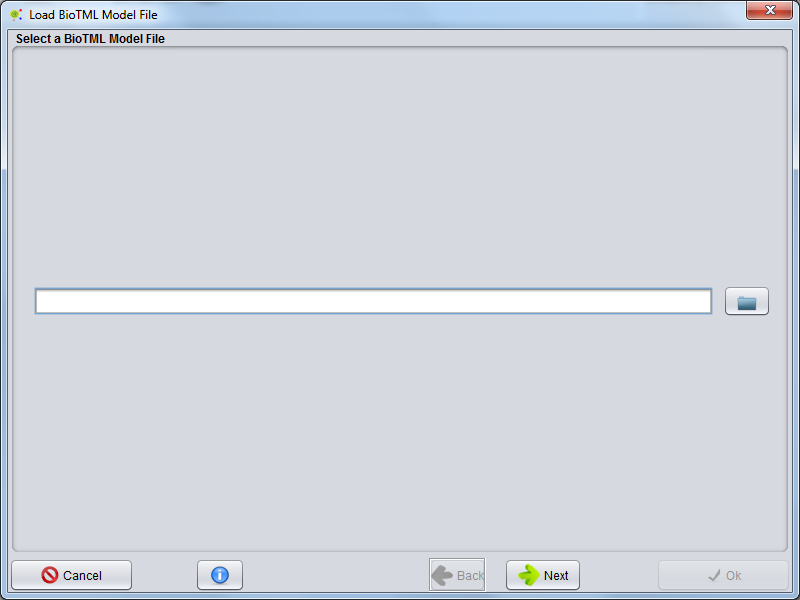
A GUI appears allowing to select an BioTML NER model file. Select a compatible BioTML model file (could be a .zip or a .gz file) and press next. The BioTML Model loading will start.
Select_BioTML_Model_Configurations_To_Annotate
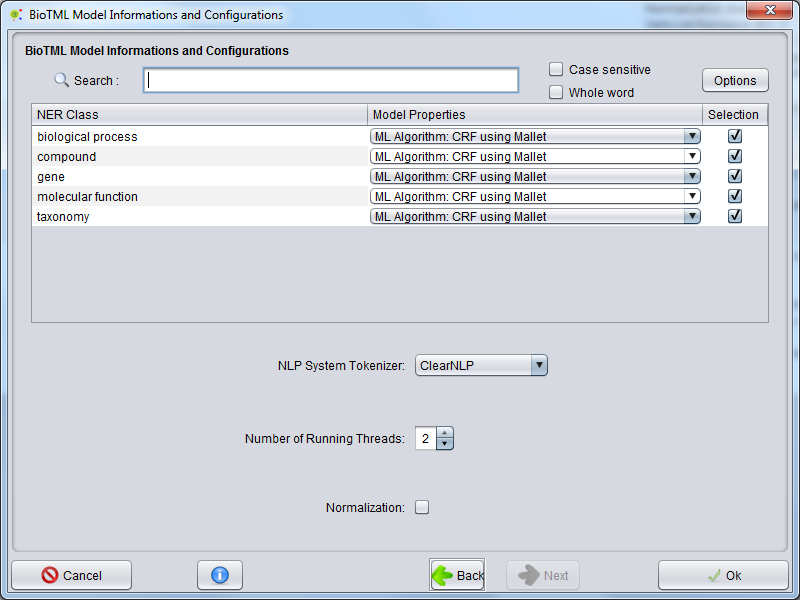
The selected model will contains NER classes to annotate and model properties that could be selected in the search table.
The NER BioTML Tagger allows you to define options for annotations, as follows:
In the NLP System Tokenizer option, you could select the natural language processing system used for tokenization of all documents in order to annotate.
In the Number of running Threads option, you can select the number of processing threads that the NER BioTML Tagger algorithm will be able to use.
In the Normalization option, you can select the @Note2 normalization system that allows to add white space between word delimiters and increase the entity recognition accuracy. This option changes the original offsets of the text..
The configuration is now complete once you press Ok. At all times you can cancel the operation clicking on the respective button.
Performing the operation
NER BioTML Tagger operation will now start and a progress window will appear, indicating the execution status of the operation. The NER BioTML Tagger operation will take a few minutes or hours depending on the number of documents, document size, number of given threads, number of model classes in the model and features used automatically by BioTML model.
When the process ends, a new NER Process object will be added to the clipboard and can be visualized through the Corpus Process View.